
Rapid Evolution of Citrate Utilization by Escherichia coli by Direct Selection Requires citT and dctA | Journal of Bacteriology
Direct selection of E. coli in minimal M9C yielded Cit mutants for both... | Download Scientific Diagram
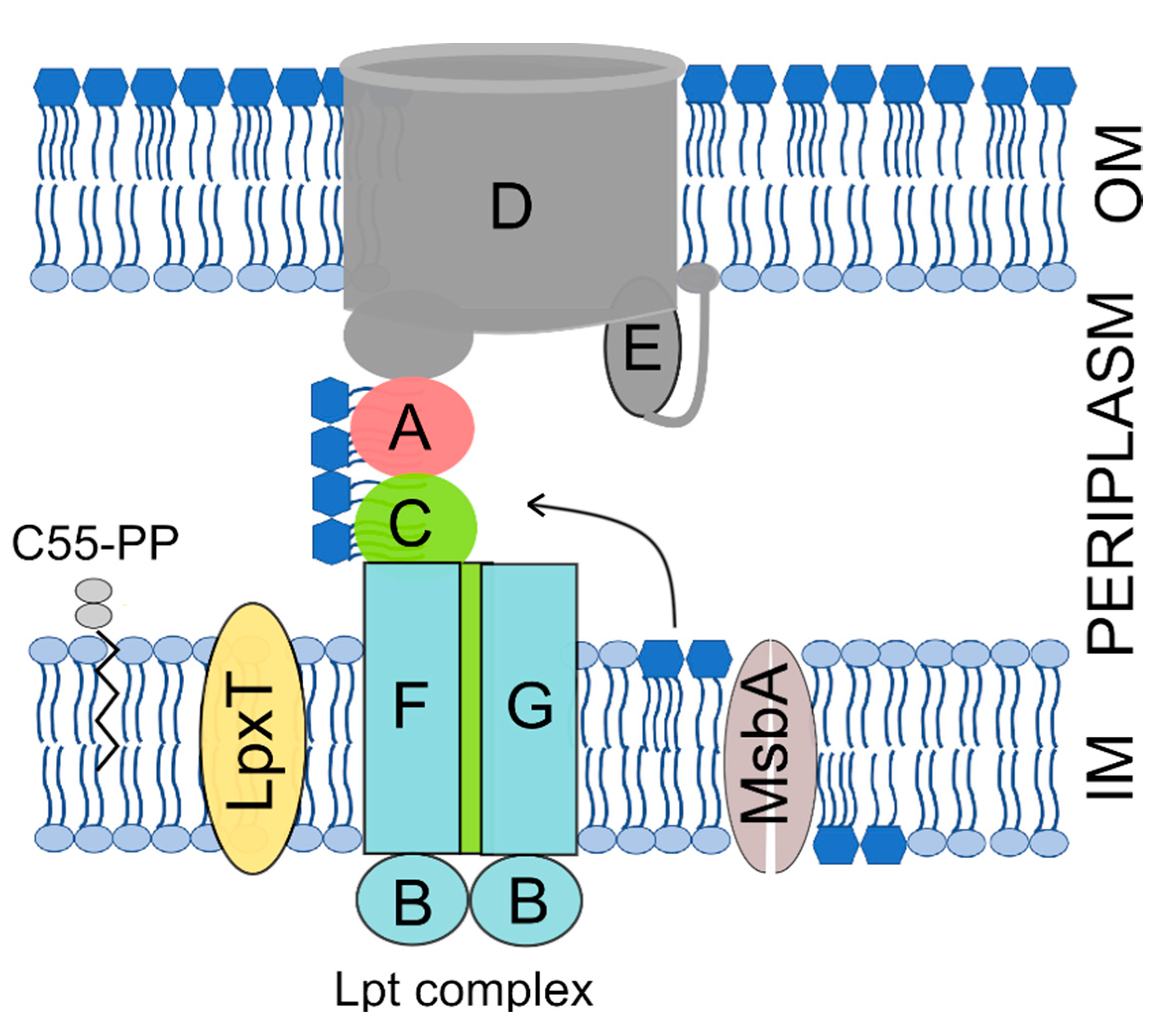
Microorganisms | Free Full-Text | Overexpression of lpxT Gene in Escherichia coli Inhibits Cell Division and Causes Envelope Defects without Changing the Overall Phosphorylation Level of Lipid A

Mutant strains of Escherichia coli lacking global regulators, arcA and fis, demonstrate better growth fitness by pathway reprogramming under acetate metabolism | bioRxiv

Protocol to identify the core gene supported by an essential gene in E. coli bacteria using a genome-wide suppressor screen - ScienceDirect

Rapid Evolution of Citrate Utilization by Escherichia coli by Direct Selection Requires citT and dctA | Journal of Bacteriology

Rapid Evolution of Citrate Utilization by Escherichia coli by Direct Selection Requires citT and dctA | Journal of Bacteriology

Escherichia coli Data-Driven Strain Design Using Aggregated Adaptive Laboratory Evolution Mutational Data | ACS Synthetic Biology

Environmental complexity is more important than mutation in driving the evolution of latent novel traits in E. coli | Nature Communications
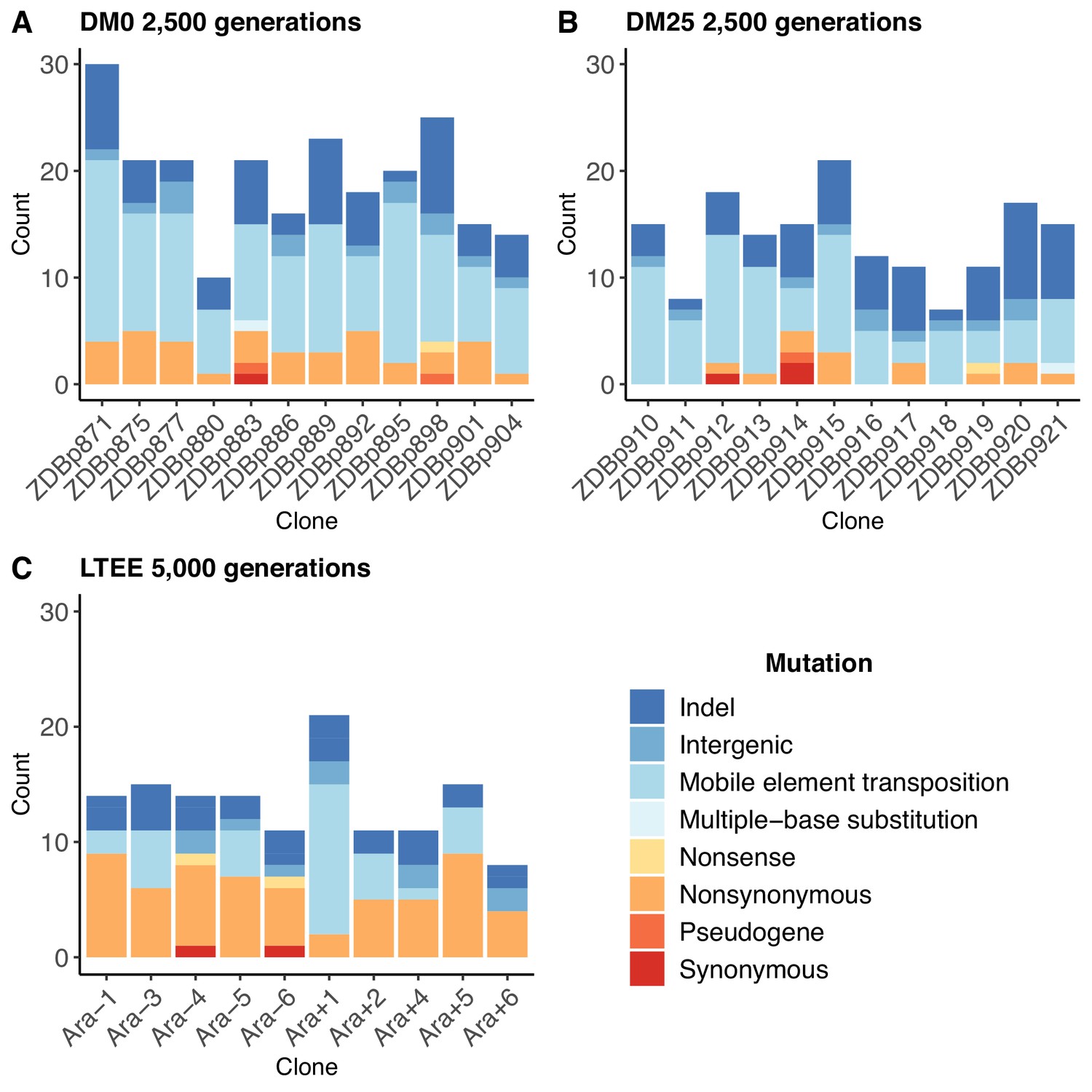
Genomic and phenotypic evolution of Escherichia coli in a novel citrate-only resource environment | eLife
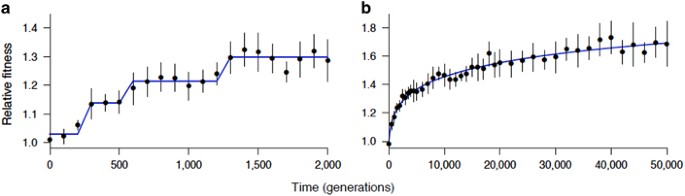
Experimental evolution and the dynamics of adaptation and genome evolution in microbial populations | The ISME Journal

Rapid Evolution of Citrate Utilization by Escherichia coli by Direct Selection Requires citT and dctA | Journal of Bacteriology
Innovation in an E. coli evolution experiment is contingent on maintaining adaptive potential until competition subsides | PLOS Genetics

Escherichia coli Data-Driven Strain Design Using Aggregated Adaptive Laboratory Evolution Mutational Data | ACS Synthetic Biology

Genomewide phenotypic analysis of growth, cell morphogenesis, and cell cycle events in Escherichia coli | Molecular Systems Biology
Innovation in an E. coli evolution experiment is contingent on maintaining adaptive potential until competition subsides | PLOS Genetics

Metabolic engineering of Escherichia coli W3110 to produce L‐malate - Dong - 2017 - Biotechnology and Bioengineering - Wiley Online Library

Santa Fe Institute on Twitter: "On @RELenski's Long-Term Evolution Experiment (#LTEE) — looking back thousands of #Ecoli generations, researchers found precursor "scaffolding" mutations that permitted later major metabolic innovations but were themselves

Recursive genomewide recombination and sequencing reveals a key refinement step in the evolution of a metabolic innovation in Escherichia coli | PNAS







